Introduction to raster data in R
Contributors: Clark S. Rushing
This activity will introduce you to working with raster data in R.
R Skill Level: Intermediate - this activity assumes you have a working knowledge of R
Need to brush up on syntax and data classes in R? See R basics for a refresher.
Objectives & Goals
Upon completion of this activity, you will:- know how to create and write rasters
- know how to do basic calculations
Required packages
To complete the following activity you will need the following packages installed: raster sp rgeosInstalling the packages
If installing these packages for the first time consider addingdependencies=TRUE
install.packages("raster",dependencies = TRUE)install.packages("rgdal",dependencies = TRUE)install.packages("rgeos",dependencies = TRUE)install.packages("sp",dependencies = TRUE)Download R script Last modified: 2019-09-20 18:26:28
What is a raster?
A raster is a spatially explicit matrix or grid where each cell represents a geographic location. Each cell represents a pixel on a surface. The size of each pixel defines the resolution or res of raster. The smaller the pixel size the finer the spatial resolution. The extent or spatial coverage of a raster is defined by the minima and maxima for both x and y coordinates.
Rasters can be created from the following data classes:
Raster formats
Raster data are stored in a variety of formats. The table below shows several commonly encountered file types. Use raster::writeFormats() to see the full list.
## Name Long Name File Extension
## 1 raster R-raster .grd
## 5 BIL Band by Line .bil
## 8 ascii Arc ASCII .asc
## 22 GTiff GeoTIFF .tiff
## 37 netCDF Network Common Data Format .nc
Creating & writing rasters
Raster data in R
Let’s begin by creating a raster from scratch. We’ll use the raster package to make an empty raster, set the extent and resolution (res) and assign values. Once we create a raster in R - we’ll take a closer look at the metadata and structure of rasters in R.
load the raster package if you haven’t already done so. If you need to install the raster package - see how to do that here
# load library
library(raster)
Now that the raster library is loaded we can use the raster() function to create a raster in R.
# Create a raster from scratch using raster
firstRaster <- raster(xmn = -100, # set minimum x coordinate
xmx = -60, # set maximum x coordinate
ymn = 25, # set minimum y coordinate
ymx = 50, # set maximum y coordinate
res = c(1,1)) # resolution in c(x,y) direction
Here is what that raster looks like in R
# Take a look at what the raster looks like
firstRaster
## class : RasterLayer
## dimensions : 25, 40, 1000 (nrow, ncol, ncell)
## resolution : 1, 1 (x, y)
## extent : -100, -60, 25, 50 (xmin, xmax, ymin, ymax)
## crs : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
Notice that the object is of class: RasterLayer has 25 rows, 40 columns and 1000 cells. The resolution (res) is 1x1 degree. The raster’s extent ranges from -100 to -60 degrees longitude and 25 to 50 degrees latitude. The coordinate reference is WGS84 by default because raster recognized our inputs as degrees longitude/latitude. It doesn’t always do so.
Setting raster values
Currently there are no values associated with the raster layer we just created. It’s empty. We can assign values to the raster in a few ways. We’ll set the values of the raster using the [] convention. See the setValues function in the raster package for another way to set values of a raster. We’ll sequence values from 1 to the number of cells within the raster. You can extract the number of cells within a raster using the ncell function.
note - the number of values you supply needs to be equivilant to the number of cells in the raster. You can however provide NA values.
# Assign values to raster
firstRaster[] <- seq(from = 1, to = ncell(firstRaster),by = 1)
# Take a look at the raster now
firstRaster
## class : RasterLayer
## dimensions : 25, 40, 1000 (nrow, ncol, ncell)
## resolution : 1, 1 (x, y)
## extent : -100, -60, 25, 50 (xmin, xmax, ymin, ymax)
## crs : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## source : memory
## names : layer
## values : 1, 1000 (min, max)
You should now notice there are a few new attributes to our raster object. We gained data source:, names and values fields. The data source attribute tells us that the raster information is stored in our memory. The names field gave a name to the values we provided. The values we supplied are now contained in the values field.
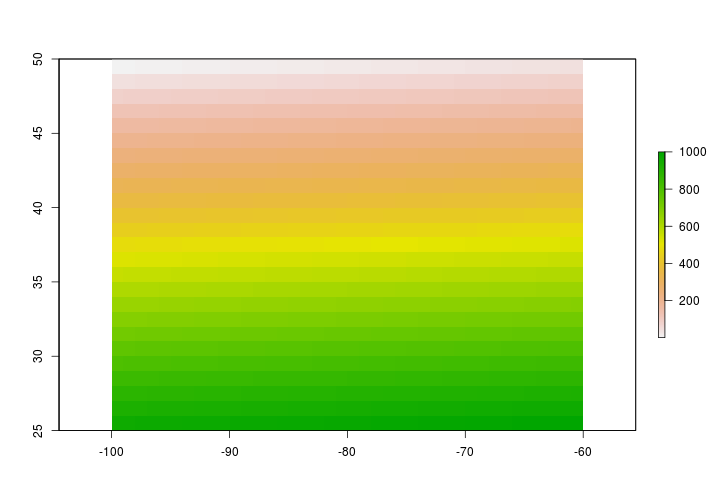
Now that the raster has values we can do a few things - like plot the raster.
plot(firstRaster)
Reading rasters from file
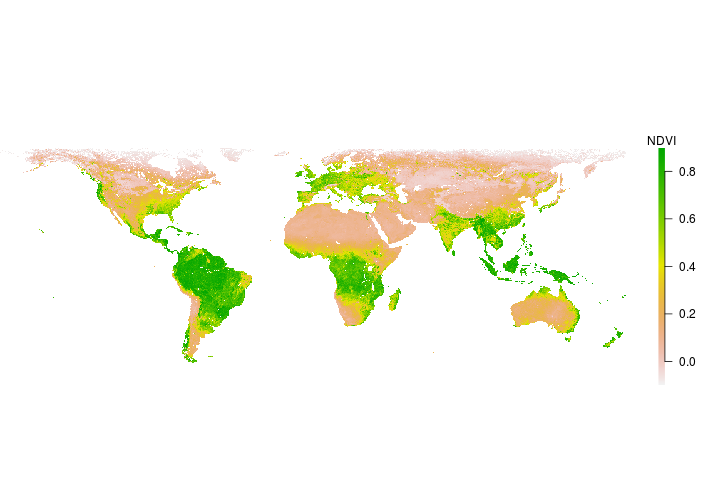
The raster() function within the raster package can also be used to read in a raster from file. Let’s read in some Normalized Difference Vegetation Index (NDVI) data from NASA’s NEO. See where to get data for other potential data sources.
# read in raster layer using raster function
# NDVI <- raster("path/to/raster/file")
NDVI <- raster::raster("../Spatial_Layers/MOD_NDVI_M_2018-01-01_rgb_3600x1800.FLOAT.TIFF")
Let’s take a look at the data
NDVI
## class : RasterLayer
## dimensions : 1800, 3600, 6480000 (nrow, ncol, ncell)
## resolution : 0.1, 0.1 (x, y)
## extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
## crs : +proj=longlat +datum=WGS84 +no_defs +ellps=WGS84 +towgs84=0,0,0
## source : /home/travis/build/mhallwor/mhallwor.github.io/Spatial_Layers/MOD_NDVI_M_2018-01-01_rgb_3600x1800.FLOAT.TIFF
## names : MOD_NDVI_M_2018.01.01_rgb_3600x1800.FLOAT
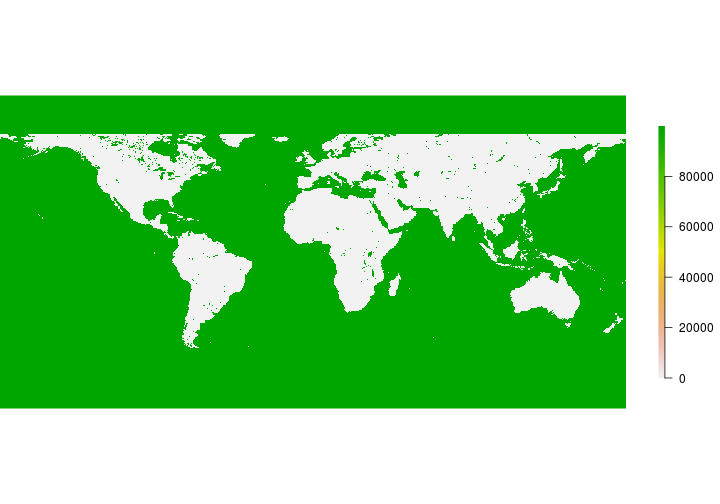
Challenge
Does the NDVI appear as you expected?